| Pasteurellaceae | |
|---|---|
| |
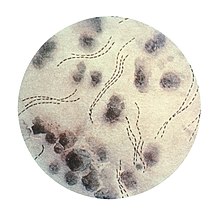
| Haemophilus ducreyi | |
| Scientific classification | |
| Domain: | Bacteria |
| Phylum: | Pseudomonadota |
| Class: | Gammaproteobacteria |
| Order: | Pasteurellales Garrity et al. 2005 |
| Family: | Pasteurellaceae Castellani & Chalmers, 1919 |
| Genera | |
|
Actinobacillus | |
The Pasteurellaceae comprise a large family of Gram-negative bacteria. Most members live as commensals on mucosal surfaces of birds and mammals, especially in the upper respiratory tract.[3] Pasteurellaceae are typically rod-shaped, and are a notable group of facultative anaerobes. Their biochemical characteristics can be distinguished from the related Enterobacteriaceae by the presence of oxidase, and from most other similar bacteria by the absence of flagella.
Bacteria in the family Pasteurellaceae have been classified into a number of genera based on metabolic properties, but these classifications are not generally accurate reflections of the evolutionary relationships between different species. Haemophilus influenzae was the first organism to have its genome sequenced and has been studied intensively by genetic and molecular methodologies. The genus Haemophilus is a notorious human pathogen associated with bacteremia, pneumonia, meningitis and chancroid. Other pathogenic members of the family Pasteurellaceae include Aggregatibacter, Mannheimia, Pasteurella, and Actinobacillus species.
Molecular signatures and phylogenetic position
[edit]Comparative analyses of Pasteurellaceae genomes have identified large numbers (>20) of conserved signature indels (CSIs) in different important proteins that are uniquely shared by all sequenced Pasteurellaceae species/strains, but are not found in any other bacteria. Based upon many other CSIs that are specific for subgroups of Pasteurellaceae species, it has been proposed to divide the family into at least two clades .[4] One proposed clade includes Aggregatibacter, Pasteurella, Actinobacillus succinogenes, Haemophilus influenzae, Haemophilus somnus, and Mannheimia succiniciproducens, while the other includes Actinobacillus minor, Actinobacillus pleuropneumoniae, Haemophilus ducreyi, Haemophilus parasuis, and Mannheimia haemolytica.
Molecular signatures in the form of CSIs have also been used to help resolve polyphyletic distribution of three main genera within the family Pasteurellaceae: Actinobacillus, Haemophilus, and Pasteurella.[5][6][7] These genera demonstrate extensive polyphyly across the family, however, CSIs have been found to be consistently shared by certain species that form a monophyletic group within each respective genus.[5] The distribution of CSIs corresponds to sensu stricto clades of "true" Actinobacillus, Haemophilus, and Pasteurella species, respectively. Since they are indicative of common ancestry, it has been postulated that the CSI distribution can be used to determine genus identity, where the species that do not share the CSI may be reclassified as a different genus. CSIs have also been found that are specific for Aggregatibacter and Mannheimia, two clinically relevant genera.[5]
The Pasteurellales, along with Enterobacterales, are of the most recently divergent orders within the Gammaproteobacteria.[8] Their distinction from all other orders are supported by the presence of several conserved signature proteins (CSPs) that are shared by these two orders, and absent from all other bacteria.[8] Pasteurellales also share additional CSPs with Enterobacterales, Vibrionales, Aeromonadales, and Alteromonadales, adding additional resolution to their evolutionary branching and phylogenetic position among the large Gammaproteobacteria class.[8]
Etymology
[edit]This taxonomical group of bacteria was named after Louis Pasteur, a name formally accepted in 1981. The designation was conceived to accommodate a collection of gram-negative organisms currently representing 34 genera and 105 species, described as “specialized commensals, primarily and potential pathogens of vertebrates–mainly mammals and birds.” [9]
References
[edit]- ^ Parker, Charles Thomas; Garrity, George M. (2018). Parker, Charles Thomas; Garrity, George M (eds.). "Taxonomy of the genus Caviibacterium Adhikary et al. 2018". doi:10.1601/tx.31260 (inactive 1 November 2024).
{{cite journal}}: Cite journal requires|journal=(help)CS1 maint: DOI inactive as of November 2024 (link) - ^ Parker, Charles Thomas; Garrity, George M. (2018). Parker, Charles Thomas; Garrity, George M (eds.). "Taxonomy of the genus Conservatibacter Adhikary et al. 2018". doi:10.1601/tx.31262 (inactive 1 November 2024).
{{cite journal}}: Cite journal requires|journal=(help)CS1 maint: DOI inactive as of November 2024 (link) - ^ Kokotovic, Branko; Friis, Niels F; Ahrens, Peter (2007). "Mycoplasma alkalescens demonstrated in bronchoalveolar lavage of cattle in Denmark". Acta Veterinaria Scandinavica. 49 (1): 2. doi:10.1186/1751-0147-49-2. ISSN 1751-0147. PMC 1766361. PMID 17204146.
- ^ Naushad, HS; Gupta, RS (Jan 2012). "Molecular signatures (conserved indels) in protein sequences that are specific for the order Pasteurellales and distinguish two of its main clades". Antonie van Leeuwenhoek. 101 (1): 105–24. doi:10.1007/s10482-011-9628-4. PMID 21830122. S2CID 15114511.
- ^ a b c Naushad S, Adeolu M, Goel N, Khadka B, Al-Dahwi A, Gupta RS (2015). "Phylogenomic and molecular demarcation of the core members of the polyphyletic pasteurellaceae genera actinobacillus, haemophilus, and pasteurella". Int J Genom. 2015: 198560. doi:10.1155/2015/198560. PMC 4363679. PMID 25821780.
- ^ Olsen I, Dewhirst FE, Paster BJ, Busse HJ (2005) Family I. Pasteurellaceae. In: Bergey’s Manual of Systematic Bacteriology, edn. 2, vol 2, pp. 851–856. Eds Brenner D. J., Krieg N. R., Garrity G. M., Staley J. T. Springer-: New York.
- ^ Dewhirst FE, Paster BJ, Olsen I, Fraser GJ (1992). "Phylogeny of 54 representative strains of species in the family Pasteurellaceae as determined by comparison of 16S rRNA sequences". J Bacteriol. 174 (6): 2002–2013. doi:10.1128/jb.174.6.2002-2013.1992. PMC 205807. PMID 1548238.
- ^ a b c Gao B, Mohan R, Gupta RS (2009). "Phylogenomics and protein signatures elucidating the evolutionary relationships among the Gammaproteobacteria". Int J Syst Evol Microbiol. 59 (Pt 2): 234–247. doi:10.1099/ijs.0.002741-0. PMID 19196760.
- ^ Partin, Clyde (October 2024). "Etymologia - Pasteurellaceae [pas′′-tər-ə-lā′-sē-ī]". Emerg Infect Dis. 30 (10). doi:10.3201/eid3010.240735. PMC 11431925. Retrieved October 2, 2024.
cites public domain text from the CDC